Good afternoon everyone!
We are trying to produce at CRI both Bst-LF and M-mmlv proteins. The results on the SDS page seems a bit weird. We wanted to upload them so you may help us troubleshooting what’s going on, and hopefully that could serve for solving doubst of future people trying this protocol.
The protocol we used: Recombinant expression and purification of codon-optimized M-MLV and Mashup
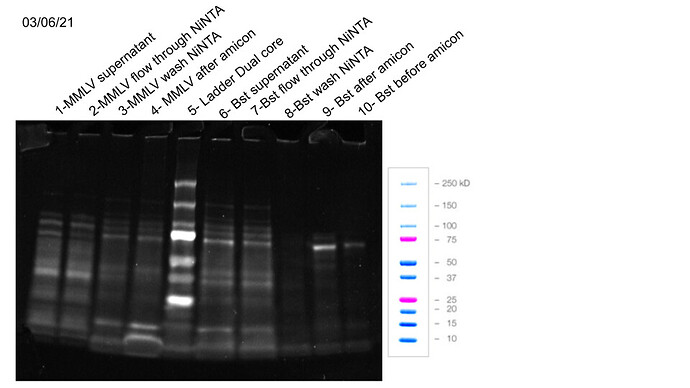
SDS page results
m-mlv questions / problems
Are we losing protein in the NiNTA flowthrough or is other 60kDa proteins?
It seems that we are loosing protein in the NiNTA wash right?
Should we incubate ON with DTT for removing the sulfur corsslinked species?
Bst questions
Are we loosing protein in the flowtrough or they are other 60kDa e.coli proteins?
Thank you in advance!
Fran
Hi there!
Sorry, just recently became aware of these comments on our protocol. I’m also sorry that the purification is so far not working for you. Lastly, I am sorry that I will have to split my answer in three messages, but I cannot upload more than one picture per message.
First of all, it does not seem that there is a lot of overexpression for either MMLV or BstLF, which is somewhat different from what we typically obtain. For example, below is a MMLV purification we did last year:
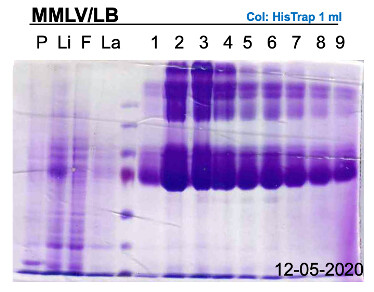
Do note that the MW of MMLV is about 70-75 kDA on the SDS-PAGE.
Typically, we obtain about 4 mg MMLV and 10 mg BstLF per liter of cell culture. It is worth noting that we observed that the extent of MMLV present in the supernatant heavily depends on the use of lysozyme before sonication, barely obtaining less than 0.5 mg of pure MMLV when we did not use lysozyme.
I also see a lot of contaminants <60 kDA in the steps after NiNTA purification. We have only seen this occurring once, when one of us mistakenly employed buffer B (which has 150 mM imidazole) instead of buffer A (which has 40 mM imidazole) for washing out proteins non-specifically bound to the NiNTA column before eluting MMLV. See the figure below (from left to right we have ladder/pellet/supernatant/flow-through/wash with B instead of A/elutions)
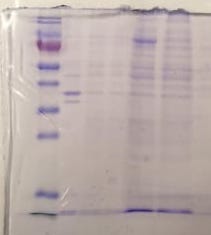
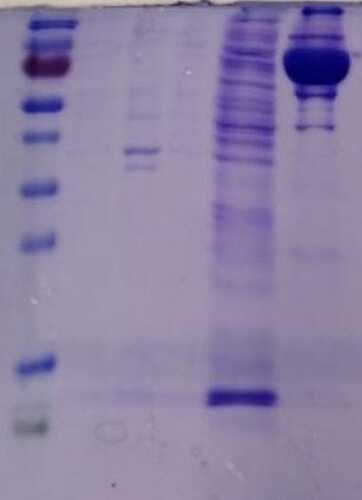
When we repeated the purification, but now correctly washing with buffer A, our results were great. The following is a purification from a few months ago (again, from left to right we have ladder/pellet/supernatant/flow-through/wash with A/elution)
Lastly, regarding the use of DTT, the reducing agent is only for eliminating higher-order aggregates (>140 kDa) that we observed when MMLV was stored without DTT. To my understanding, incubation overnight with DTT would not solve the problems that you are indicating here (i.e. low protein over expression, lack of purified protein).
And this is my third and last message! Hope it helps!
All the best,
Cesar
Hi Cesar and thank you for such a detailed and amazing answer,
We definitely don’t have that much overexpression, and we are trying to troubleshoot that. We are trying an autoinducible medium right now. We don’t know if it’s the bacteria or the IPTG concentration, we are double-checking everything right now but it seems okay.
Thank you also for your insights on the possible buffer/imidazole cofussion. We are retrying today the purification, so I’ll keep you posted about the results.
Gracias de nuevo!
Fran