MMLV RT
Reagent Sharers
Alex Brown
Summary
This is a RT derived from Moloney Murine Leukemia Virus containing D200N, L603W, T330P, L139P and E607K, for high processivity and thermostability (Baranauskas, 2012 PMID: 22691702)
Expression/Production Information
Alex has posted the expression/purification protocol at protocols.io
How to get the plasmid/material/reagent
Currently in Alex´s hands, OBL, FreeGenes, iBio_Chile and UBA_Argentina
This is vector received from Alex and being used in Chile.
FreeGenes is also distributing this vector.
Terms
Under openMTA from Alex, being able to redistribute under openMTA from iBio/Chile too.
References
Baranauskas, 2012 PMID: 22691702
1 Like
Right now working with MMLV, everything has worked great!
We are optimizing the conditions of use, but it seems that it is an enzyme with very robust activity in different conditions.
2 Likes
We are also working on the expression of MMLV in Buenos Aires. Everything goes fine so far, but we have a little doubt about the protocol you shared. The lysozyme concentration on Buffer B is 1mg/ml. Isn’t high? Have someone tried without lysozyme or using a lower concentration?
1 Like
Cesar Ramirez (in Chile) said they used 0.1 mg/ml and works.
1 Like
Another purification protocol for MMLV has been added to the collection in protocols.io
https://www.protocols.io/view/recombinant-expression-and-purification-of-codon-o-bhcvj2w6
It is working for us in Chile.
Thanks to Cesar Ramirez group!
1 Like
Hey guys!
I think I saw some aggregration on my SDS-PAGE (didnt confirm by Western Blot, though).
This publication: https://pubmed.ncbi.nlm.nih.gov/19330487/ states, that MMLV tends to aggregate in Imidazol and suggests an on column-cleavage elution with high recovery rate.
I’ve also read (dont know where right now ^^), that MMLV needs to be cleaved, otherwise there is a huge decrease in acitivity.
Any thoughts on that? I will try on-column cleavage the next run I guess.
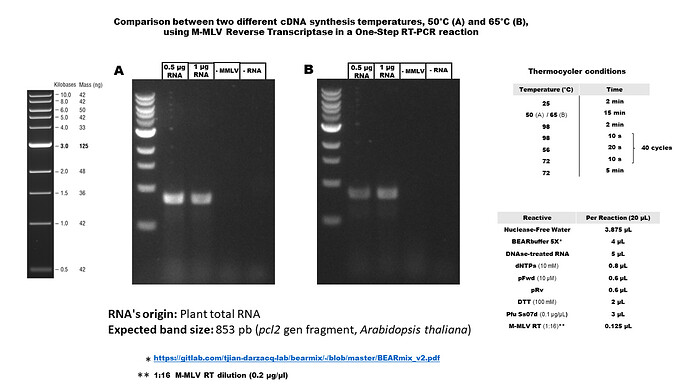
In case of any interest, we are using MMLV and Pfu Ss07d in One-Step PCR reactions. Traditionally we have used MMLV for cDNA synthesis at 50 ° C, but recently we tested at 65 ° C and it works almost the same. I attach a diagram that summarizes the conditions used.
2 Likes
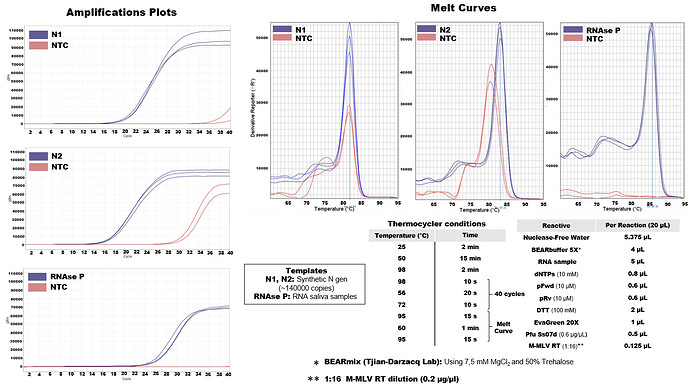
I attached the type of results that we are obtaining in the One-Step RT-qPCR using the enzymes MMLV-RT and Pfu-Ss7od. In this case, the template corresponds to the synthetic N gene of SARSCoV2. Our main problem has been the contamination of the negative controls with something that appears to be non-specific amplification of the primers (i.e: N2) and / or contamination with the synthetic template (i.e: N1). Now we are working with clinical samples and we are obtaining very good results with the conditions and thermocycler program indicated in the figure. We still have nonspecific amplification problems but the positive and negative samples are distinguishable by Tm calling analysis. There is still room for improvement, though.
Pfu-Sso7d and MMLV were purified as publicly described by us in Recombinant protein expression and purification of codon-optimized Pfu-Sso7d and Recombinant expression and purification of codon-optimized M-MLV and Mashup
1 Like
Greetings everyone 
Could you tell me if anyone has this reagent, where could I get it or from who??
Btw, I’m a student and i realy need this MMLV
All the best, hoping for a feedback 
@kostetskiijr where are you geographically based?
@jenny_molloy
Hi, right now I’m located in Ukraine, Kyiv city