Hi, we would like to share our progress on a small project with some friends of the reGOSH/GOSH community (Nano, Nico, Fran, Wladi) and the Open Bioeconomy Lab.
This project addresses a common issue in Latin America: namely “how to teach hands-on PCR in universities and schools”. It is normal to hear that students graduate in biotech/biology/vet/agronomy/medicine without ever performing a PCR. A lot of progress has been made to facilitate access to low cost thermocyclers in the last few years (GaudiLabs, Shingo, Chai PCR and the not-open but mid-cost miniPCR). However, access to reagents has remained more difficult. PCR reagents are expensive to obtain. They take a long time to arrive. Importation logistics (customs and cold-chain) are not easy and in most countries you have to get it from official distributors (adding an extra cost) (a quantitative study here from BID/Inter American Development Bank).
Producing and purifying enzymes, as many of us do in Reclone, can reduce costs by 90% (we do it routinely in our labs). However, the expertise and infrastructure required to do protein purification is very scattered and not accessible to everybody nor sustainable (mostly done for domestic use only).
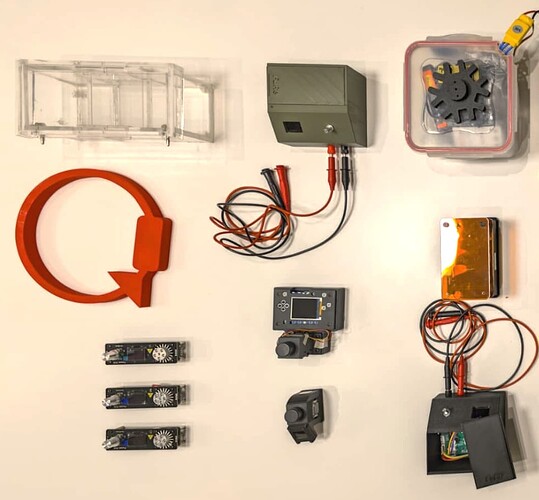
Very recently, a game-changing protocol from Andy Ellington´s lab demonstrated that PCR reactions can be prepared directly from bacteria containing the PCR enzyme inside. The PCR is performed using intact bacteria containing the enzyme, thus eliminating the need for prior lysis and purification. In other words “you pipette bacteria into the PCR reaction as if you were pipetting a pure enzyme”. However, this protocol still requires the use of chemical inducers (such as IPTG) as well as incubators and shakers to grow bacteria in liquid cultures, which is not available in high schools or most univ teaching labs. In response to this, the protocol has been further extended by the Open Bioeconomy Lab to allow the growth and harvest steps directly from solid media, petri dishes containing low cost autoinduction media (link). This is a very nice trick that avoids the use of expensive inducers and equipment for maintaining liquid cultures; just grow bacteria on plates and harvest from them directly with a clean microscope slide or a razor blade. We took it from there with Jenny/OBL and all the people mentioned above to enable the implementation of this protocol with a low cost lab ecosystem:
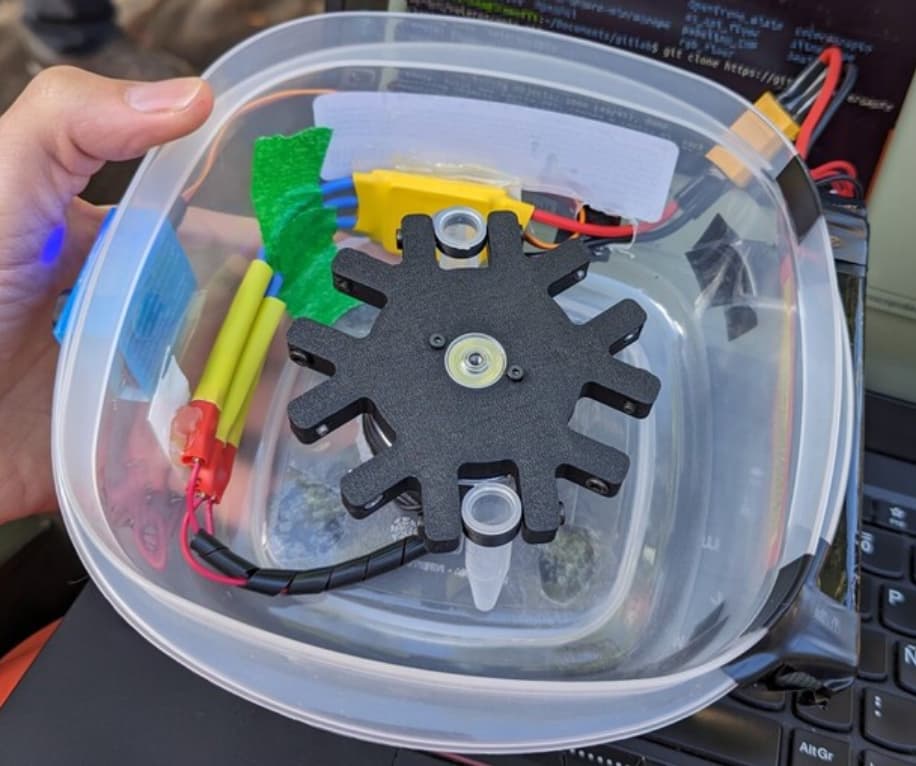
- The use of the tupperfuge for low cost cleaning of collected cells (it is required to remove pieces of agar and other “debris” from the bacteria harvesting step.
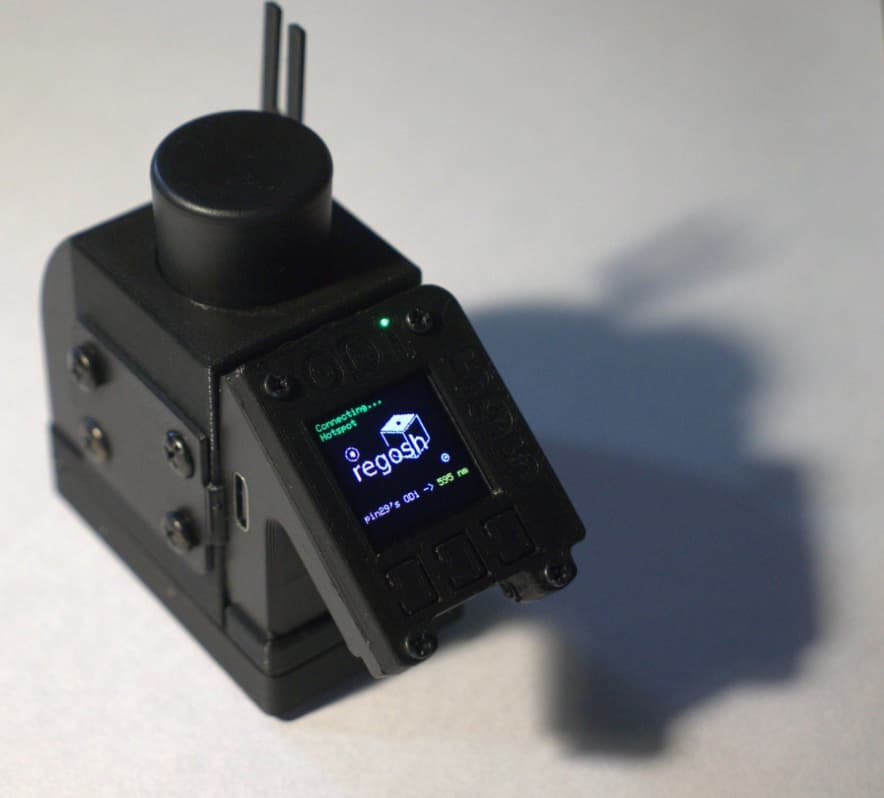
- The measurement of OD with an open colorimeter after harvesting and washing the cells. This is an important step for deciding how much of the bacterial suspension needs to be diluted before using it in a reaction.
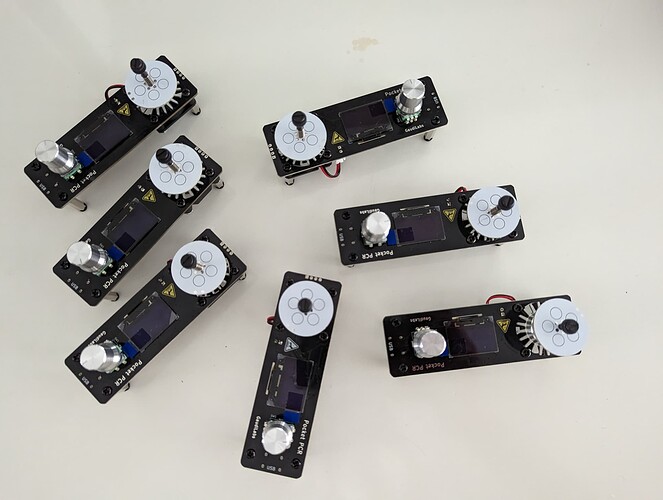
- The use of pocketPCR from GaudiLabs for the amplification reaction. We are big fans of this device. It works robustly everywhere, always (just be sure you have a good power supply to deliver the required Amp).
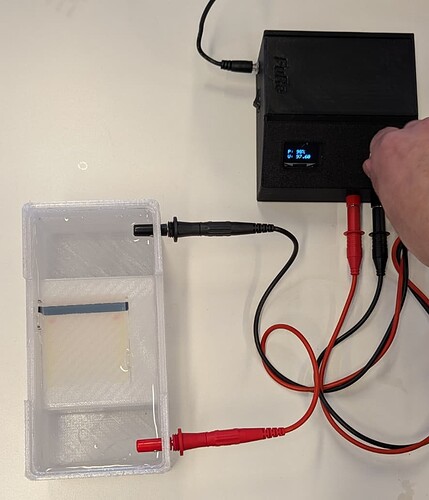
- The separation of the amplified DNA by electrophoresis in a chamber and open source power supply that can be built locally (here).
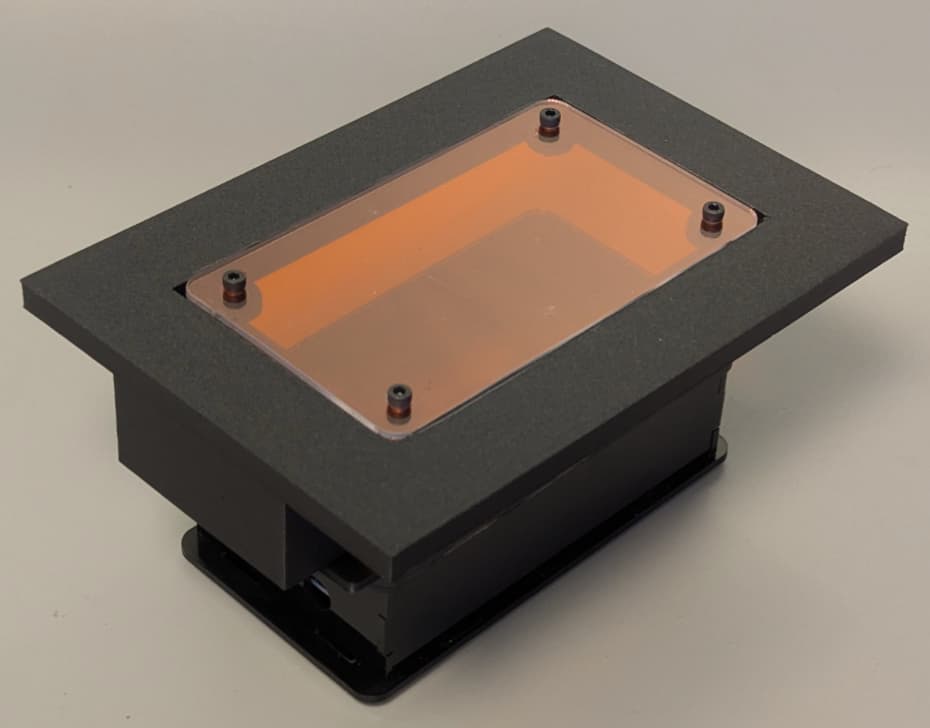
- The visualization of results with an open transilluminator, which can be built locally in Chile/Argentina without depending on imported acrylics and LEDs. A nice collaboration with Jo from IORodeo here. We have worked with Fran Quero to adapt the IOrodeo transilluminator to be more fabricable in Chile/Argentina. First, we changed the superbrightleds (details in the repo and docs linked in readme) that are very nice but expensive and need to be soldered by hand, one by one. Fran designed a new PCB with SMD blue LEDs that work very well and you can order them already mounted in the PCB (we tried many different options, see the repo if interested). Next, we replaced the acrylic filters with leefilters (used in the GMOdetective project). With all these changes, the device works really nice but at a fraction of the original cost. We also add a usb-c alternative power supply because jack barrels are sometimes mixed up in a lab (e.g. 12,V, 15V, etc) and it is easy to burn things.
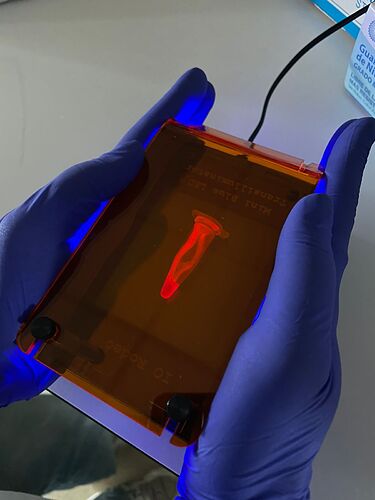
The protocol in full details is here (link) but in brief: grow E. coli containing the pTI vector (an open access vector engineered by our group for protein expression (link)) containing a polymerase named OpenVent, fused to a red fluorescent protein (so you can see red fluor or pink color with naked eye, as a proxy for checking polymerase expression). The expression of these genes is controlled by IPTG (an expensive analogue of lactose metabolites), but here is the trick from Jenny´s protocol: autoinduction media in which cells first consume glucose and then lactose, these metabolites activate gene expression. You incubate the plates at 37 overnight. We are testing bed blankets for this but any device that reaches 37C would do the job (e.g. kombucha fermenter mats, or close to an oven in winter). When the plates become pink, it means the enzyme and red fluorescent protein has expressed and you are ready to collect cells. Next, you have to collect the cells with a razor blade or microscope slide and collect them in an eppendorf. Here, you use Nico’s tupperfuge to precipitate the cells and wash any agar and debris carried during the harvesting step. Next, an open colorimeter is used to calculate the range of working bacterial densities. We are working on a method to measure red fluorescence with the transilluminator (same used below for the gel) and compare notes with OD-based measurements (sometimes bacteria growth does not equal good protein production, the two processes could be decoupled). After running the gel on electrophoretic chambers, it is visualized in a blue transilluminator.
We have tested this ecosystem in many different workshops with more than 50 teachers and we use it routinely in our university courses (>100 students in total).
Challenges ahead:
A hard one: We still rely on other reagents for PCR.
A more addressable one: how to scale up hardware production and sale for people that don’t have the time or intention to build their own
Please join the project through comments here, or in the doc (or particular issues in each repo).
Repositories (Ecosistema de Hardware Abierto para Biología Molecular – REGOSH):
- Taperfuga
- Colorimeter
repo & manual: Technical Overview - OLA is Open Lab Automata
Design inspired by:Open Colorimeter – IO Rodeo
- Pocket PCR
- Electrophoresis chamber
- Transiluminador
Funding
Funding from PME iBio Institute - National Research and Development Agency (ANID) Millennium Initiative, Fondecyt Regular 1241452 & Fondecyt Exploration 13220075
Chan Zuckerberg Initiative and Ibero-American Program for Science and Technology for Development (CYTED), network ’RELARUS – Latin American Network of Free Access Reagents for One Health’ (225RT0168).
Contributions
Cellular reagents & plate induction:Open Bioeconomy Lab
Transilluminator local version: Fran Quero, Free Technology Lab, IOWlabs, modified from original version from IORodeo with a lot of conversations with Joanne.
Gel chamber: BioFabUC, Free Technology Lab, IOWlabs,
Coloremeter: Nicolas Mendez (BE and pin29Fernando Castro (Ayllu Agroecological Laband pin29) with help from IORodeo
Gel power source:IOWlabs,
Taperfuga:Nico Mendez (BE and pin29)
PocketPCR: GaudiLabs
Millennium-PME workshops with schools: ScienceHub Punta Arenas, Austral Science Node, CADI-CEBIMA-University of Magallanes, Gudrun Kausel (UACh), Guillermo Valenzuela (USS)